Introduction¶
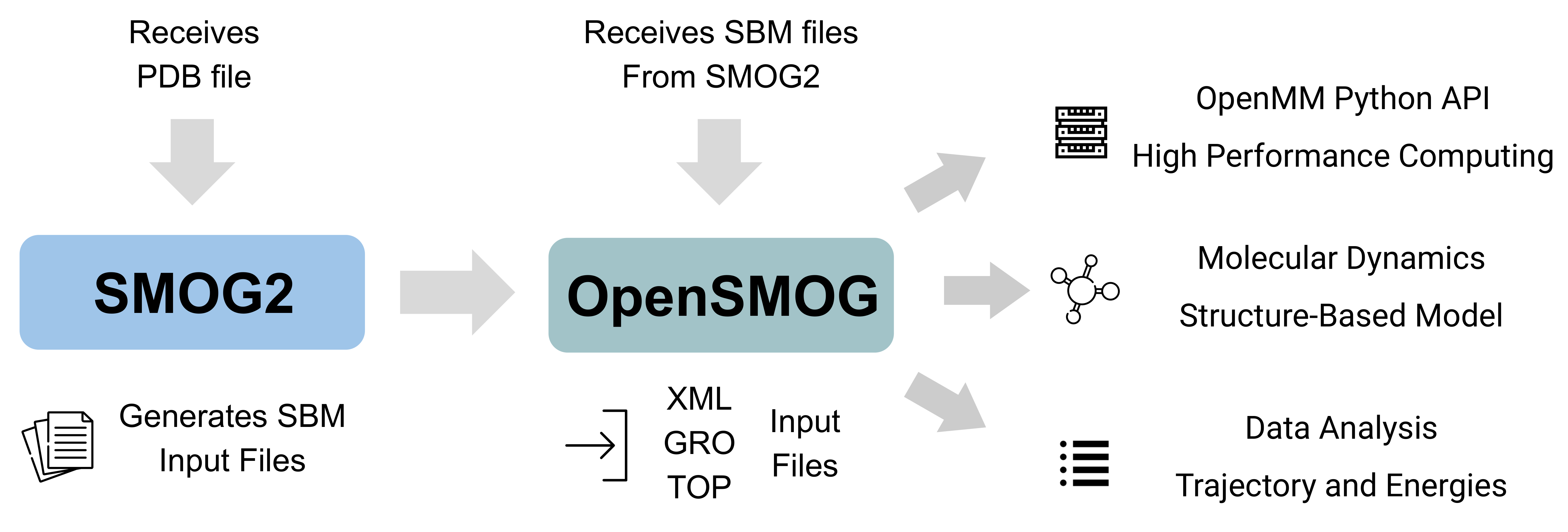
OpenSMOG is a Python library for performing molecular dynamics simulations using Structure-Based Models [1]. OpenSMOG uses the OpenMM Python API that supports a wide variety of potential forms, which includes the commonly employed C-alpha [2] and All-Atom [3] models.
The input files are generated using the SMOG2 software package with the flag -OpenSMOG. Details on SMOG2 usage can be found in the SMOG2 User Manual.
- 1
Jeffrey K Noel, Mariana Levi, Mohit Raghunathan, Heiko Lammert, Ryan L Hayes, José N Onuchic, and Paul C Whitford. Smog 2: a versatile software package for generating structure-based models. PLoS computational biology, 12(3):e1004794, 2016.
- 2
Cecilia Clementi, Hugh Nymeyer, and José Nelson Onuchic. Topological and energetic factors: what determines the structural details of the transition state ensemble and “en-route” intermediates for protein folding? an investigation for small globular proteins. Journal of molecular biology, 298(5):937–953, 2000.
- 3
Paul C Whitford, Jeffrey K Noel, Shachi Gosavi, Alexander Schug, Kevin Y Sanbonmatsu, and José N Onuchic. An all-atom structure-based potential for proteins: bridging minimal models with all-atom empirical forcefields. Proteins: Structure, Function, and Bioinformatics, 75(2):430–441, 2009.